FAF-Drugs (Free ADME-Tox Filtering Tool version 4.0) is a program for filtering large compound libraries prior to in silico screening experiments or related modeling studies. The main goal is the computational prediction of some ADME-Tox properties (Adsorption, Distribution, Metabolism, Excretion and Toxicity) in order to assist hit selection before chemical synthesis or ordering.
FAF-Drugs employs pre-defined filters, but users can also customize their own filtering parameters by using the Filter-Editor service.
FAF-Drugs3: a web server for compound property calculation and chemical library design.
Nucleic Acids Res. 2015 Jul 1;43(W1):W200-7.
The FAF-Drugs2 server: a multistep engine to prepare electronic chemical compound collections.
Bioinformatics. 2011 Jul 15;27(14):2018-20.
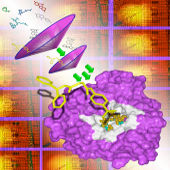
Frog is intended to generate 3D for drugs, usually described using a 1D or 2D representation. Frog performs isomer identification from ambiguous compound description. Frog is able to generate multi-conformations per isomer.
Frog2: Efficient 3D conformation ensemble generator for small compounds.
Nucleic Acids Res. 2010 Jul;38(Web Server issue):W622-7.

Pred-O3 is a web server that allows to explore interactions between molecules-odors and molecules-olfactory receptors. The server integrates a large database of odorant molecules tested on human, mouse and rat olfactory receptors. Furthermore, it allows to predict molecule-odor and molecule-olfactory receptor interactions.
Pred-O3, a web server to predict molecules, olfactory receptors and odor relationships.
Nucleic Acids Res. 2024 Jul 5;52(W1):W507-W512.

MTiOpenScreen and MTiAutoDock are open-access web services designed to support drug discovery and chemical biology by enabling small molecule docking and virtual screening. These tools facilitate the identification of new bioactive compounds through accessible, user-friendly computational workflows. By making virtual screening widely available, especially to academic researchers, they help accelerate early-stage drug discovery efforts.
MTiOpenScreen: a web server for structure-based virtual screening.
Nucleic Acids Res. 2015 Jul 1;43(W1):W448-54.
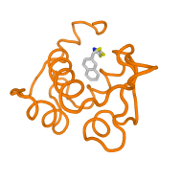
The SeamDock on-line service integrates different docking tools in a common framework that makes possible to undergo ligand global and/or local docking and a hierarchical approach combining the two for easy interaction site identification. This service does not require advanced computer knowledge and it works without installation of any programs with the exception of a common web browser.
SeamDock: An Interactive and Collaborative Online Docking Resource to Assist Small Compound Molecular Docking.
Front Mol Biosci. 2021 Sep 17;8:716466.
Docking_py, a python library for ligand protein docking.
Zenodo (2020), http://doi.org/10.5281/zenodo.4506970.
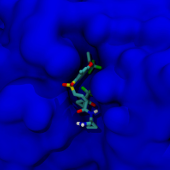
BactPepDB is a database of predicted peptides from an exhaustive survey of complete prokaryote genomes. It provides insights about candidate peptides, and provides information about their conservation, together with some of their expected biological/structural features. The BactPepDB interface allows to search for candidate peptides in the database, or to search for peptides similar to a query, according to the multiple properties predicted or related to genomic localization.
BactPepDB: a database of predicted peptides from an exhaustive survey of complete prokaryote genomes.
Database (Oxford). 2014 Nov 6;2014. Print 2014.
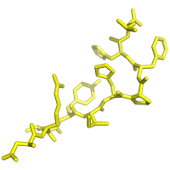
SolyPep is a fast and flexible random sequence generator for producing peptides selected for their aqueous solubility. The server first generates the required number peptides of desired length. This input library can alternatively be provided by the user in fasta format by pasting the sequences in the input field or by uploading a file. Each peptide sequence can be filtered according to a set of simple rules designed to guarantee the peptide solubility. The next step of the service will offer the processing of individual peptide sequences into 3D coordinates, files for use direct in autodock or vina can optionnally be added to the archive. It is important to note that the 3D generation step aims at providing starting conformations for flexible docking using a tool such as autodock, not predicting the conformation of the peptides in solution.
SolyPep: a fast generator of soluble peptides
in preparation.

DaReUS-Loop is a web server for the (re-)modeling of loops in homology models. It follows a data-based approach, identifying loop candidates by mining the complete set of experimental structures available in the Protein Data Bank (PDB).
Candidate loops are filtered based on the sequence and then ranked using the local conformation profile and the structural fit. DaReUS-Loop returns ten loop models for each individual loop region.
DaReUS-Loop requires an initial homology model, and can be used for either loop modeling or loop remodeling. For a single loop region, the results are identical. For multiple loop regions, remodeling typically gives more accurate results.
DaReUS-Loop: a web server to model loops in homology models.
Nucleic Acids Res. 2019 Jul 2;47(W1):W423-W428.

PEP-Cyclizer is a tool to assist the design of head-to-tail peptide cyclization, a well known strategy to enhance peptide resistance to enzymatic degradation and thus peptide bioavailability.
PEP-Cyclizer adresses two complementary features:
PEP-Cyclizer: a web server for sequence and structure prediction of peptide head-to-tail cyclization.
submitted.

PEP-FOLD is a de novo approach aimed at predicting peptide structures from amino acid sequences.
This method, based on structural alphabet SA letters to describe the conformations of four consecutive residues, couples the predicted series of SA letters to a greedy algorithm and a coarse-grained force field.
PEP-FOLD4: a pH-dependent force field for peptide structure prediction in aqueous solution.
Nucleic Acids Res. 2023 Jul 5;51(W1):W432-W437.
PEP-FOLD3: faster de novo structure prediction for linear peptides in solution and in complex.
Nucleic Acids Res. 2016 Jul 8;44(W1):W449-54.
PEP-FOLD: an updated de novo structure prediction server for both linear and disulfide bonded cyclic peptides.
Nucleic Acids Res. 2012 Jul;40(Web Server issue):W288-93.

SABBAC is an on-line service devoted to protein backbone reconstruction from alpha-carbon trace. It is based on the assembly of fragments issued from library of reduced size, resulting from the encoding of the protein trace in an HMM-derived structural alphabet. The assembly of the fragments is achieved by a greedy algorithm, using an energy based scoring inspired from the OPEP force field. Alpha-carbon coordinates remain unaffected. SABBAC simply positions the missing backbone atoms, no further refinement is performed. From our tests, SABBAC performs equal or better than other similar online approach and is robust to deviations on the alpha-carbon coordinates.
SABBAC: online Structural Alphabet-based protein BackBone reconstruction from Alpha-Carbon trace.
Nucleic Acids Res. 2006 Jul 1;34(Web Server issue):W147-51.
SA-Frag is a service that will, given an amino acid sequence, return 3D fragments predicted to match the various positions of the sequence. SA-Frag will thus return an alignement of the fragments identified with the query and a collection of 3D structures corresponding to the fragments in the PDB format.
Detecting protein candidate fragments using a structural alphabet profile comparison approach.
PLoS One. 2013 Nov 26;8(11)
SAFlex is a structural alphabet (SA ) that allows the protein tertiary structure backbones to be encoded into a sequence of short dependent structural blocks. Each structural block is called structural letter (SL) and is composed of four amino acid residues. SAFlex consists mainly of 27 Structural letters. These letters are the representative recurrent short structural 3D building blocks, identified by the Hidden Markov Model (HMM).

pepATTRACT is a fully blind protein–peptide docking protocol that requires no prior knowledge of the binding site, yet matches or outperforms local docking methods. The web server version runs the rigid-body stage in about 10 minutes, making it fast and scalable. Its blind nature and speed make it ideal for large-scale, proteome-wide docking studies.
De Vries SJ, Rey J, Schindler CEM, Zacharias M, Tuffery P.
The pepATTRACT web server for blind, large-scale peptide-protein docking.
Nucleic Acids Res. 2017 Apr 29.

PEP-SiteFinder is a service aimed at identifying patches on a protein surface, which a peptide of specified sequence is likely to interact with.
PEP-SiteFinder: a tool for the blind indentification of peptide binding sites on protein surfaces.
Nucleic Acids Res. 2014 May 6.
HHalign-Kbest is useful to automatically obtain optimized alignments and models in case of low sequence identity (<35%) between a query and a template protein. It can generate k suboptimal (e.g. top-k scoring) alignments rather than only the optimal one which may contain small to large errors.
HHalign-KBest: exploring sub-optimal alignments for remote homology comparative modeling
Bioinformatics. 2015 Dec 1;31(23):3850-2.

DockSurf is a molecular docking tool designed to predict and explore protein–surface interactions, a key challenge in bionanotechnology. It treats protein/inorganic surface modeling as a docking problem and generates a map of possible conformations using Euler angle sampling. Available on the RPBS platform, DockSurf provides fast, accessible insights into protein–surface interfaces that are often difficult to study experimentally.
DockSurf: A Molecular Modeling Software for the Prediction of Protein/Surface Adhesion.
J Chem Inf Model. 2023 Aug 28;63(16):5220-5231.

ColabFold is a structure prediction service deployed within our JupyterHub interface, allowing users to run AlphaFold2 directly from their web browser. The service is backed by our HPC infrastructure and includes a locally hosted MMseqs2 server, enabling efficient and secure computation without external dependencies. It is particularly suited for academic users needing reliable 3D structure predictions of proteins or protein complexes.
The service is backed by our HPC infrastructure and includes a locally hosted MMseqs2 server, enabling efficient and secure computation without external dependencies. It is particularly suited for academic users needing reliable 3D structure predictions of proteins or protein complexes.
ColabFold: making protein folding accessible to all.
Nat Methods. 2022 Jun;19(6):679-682.

InterEvDock is a server for protein docking running the InterEvScore potential specifically designed to integrate evolutionary information in the docking process. The InterEvScore potential was developed for heteromeric protein interfaces and combines a residue-based multi-body statistical potential with evolutionary information derived from the multiple sequence alignments of each partner in the complex.
InterEvDock3: a combined template-based and free docking server with increased performance through explicit modeling of complex homologs and integration of covariation-based contact maps.
Nucleic Acids Res. 2021 Jul 2;49(W1):W277-W284.
InterEvDock2: an expanded server for protein docking using evolutionary and biological information from homology models and multimeric inputs.
Nucleic Acids Res. 2018 Jul 2;46(W1):W408-W416.

Proteo3Dnet is a web server dedicated to the analysis of mass spectrometry interactomics experiments. Given a flat list of proteins, its aim is to organize it in terms of structural interactions to provide a clearer overview of the data. This is achieved using three means:
Proteo3Dnet: a web server for the integration of structural information with interactomics data
Nucleic Acids Res. 2021 Jul 2;49(W1):W567-W572.
BCSearch is a fast and flexible approach to identify linear fragments similar to a query in large collections of structures. It addresses two basic questions:
BCSearch is based on a new similarity approach, based on a Binet Cauchy (BC) kernel. The approach measures the correlation between the volumes of all the tetraedron of the query and that of a target. The similarity (BCscore) is scored between -1 and 1, where a value of 1 corresponds to the exact same conformation than the query, and -1 to the mirror conformation. Values close to 0 correspond to unrelated fragments. The BCscore is more stringent than other criteria such as the alpha carbon RMS deviation. Particularly, fragments with partly dissimilar shapes are poorly scored and consequently collections of matches are usually less noisy, which makes them better suited for the analysis of the local structure-sequence relationship. In addition, since no superimposition is required, the similarity search is very fast, making possible to mine large collections of structures.
BCSearch: fast structural fragment mining over large collections of protein structures.
Nucleic Acids Res. 2015 Jul 1;43(W1):W378-82.

PatchSearch allows local nonsequential searching for similar regions, called patches, on protein surfaces. It is based on detection of quasi-cliques in product graphs representing all the possible matchings between a patch and compared structures.
PatchSearch: a web server for off-target protein identification.
Nucleic Acids Res. 2019 Jul 2;47(W1):W365-W372.
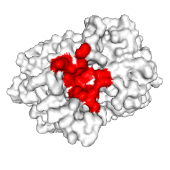
wwLigCSRre is intended for searching banks for compounds similar to a query, based on both coordinates and physico-chemical properties of atoms.
wwLigCSRre: a 3D ligand-based server for hit identification and optimization.
Nucleic Acids Res. 2009 Jul;37(Web Server issue):W504-9.
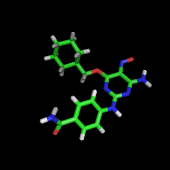
Yakusa (Yet Another K-Uples Structure Analyser) is a program devised to rapidly scan a structural database with a query protein structure. It searches for the longest common substructures, called SHSP for "structural high scoring pairs'', between a query structure and every structure in the structural database. It makes use of protein backbone internal coordinates (α angles) in order to describe protein structures as sequences of symbols. It uses a deterministic finite automaton for pattern matching.
YAKUSA: a fast structural database scanning method.
Proteins. 2005 Oct 1;61(1):137-51.

fpocket is a very fast, open source protein pocket (cavity) detection algorithm based on Voronoi tessellation. It was developed in the C programming language and is currently available as command line driven program, and from now as a web server too.
Since its first release, the new package mdpocket has been developped. This package has been designed to track cavities on different, aligned protein structures, and thus can be used to:
Currently available as a web server only, mdpocket will be released in the fpocket official distribution as soon as the corresponding scientific paper will be published.
fpocket: online tools for protein ensemble pocket detection and tracking.
Nucleic Acids Res. 2010 Jul;38(Web Server issue):W582-9.
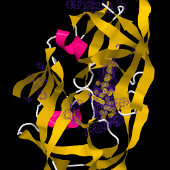
iSuperpose performs the 3D superposition of protein structures by best superimposing the alpha-carbons (or the backbone) of the proteins given a alignment specifying the correspondence between the structures. If no alignment is provided, a structural alignment will be calculated using TMalign. One the alignement is identified, the superposition is achieved using a quaternion based procedure using a specific eigen value calculation implementation. See QBestFit.
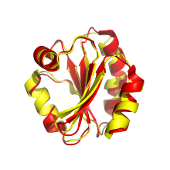
MIR is a program that allows to determine residues involved in the core of proteins. A Monte Carlo algorithm is used to simulate the early steps of protein folding and the mean number of neighbours is calculated after 10 steps. Residues surrounded by many others may play a role in the compactness of the protein and thus are called Most Interacting Residues (MIR).
Protein intrachain contact prediction with most interacting residues (MIR).
Bio-Algorithms and Med-Systems 2014 Nov 27;10(4):227-242
PockDrug is a web tool for predicting the druggability of protein pockets — their ability to bind drug-like molecules with high affinity. Unlike many models limited to a single pocket estimation method, PockDrug is robust across various estimation strategies, whether ligand-guided or not. It handles both holo and challenging apo protein structures, offering consistent and accurate predictions even in the presence of pocket boundary uncertainties. PockDrug outperforms recent models, particularly for apo structures, and supports both user-defined and server-estimated pockets.
PockDrug-Server: a new web server for predicting pocket druggability on holo and apo proteins.
Nucleic Acids Res. 2015 Jul 1;43(W1):W436-42.

PCE performs the calculation of the electrostatic potentials for a protein by solving numerically the Poisson-Boltzmann equation (the Finite Difference Poisson- Boltzmann method, FDPB). It is a server adaptation of the MEAD potential program (MEAD: Macroscopic Electrostatics with Atomic Detail, D. Bashford).
Two types of services are currently proposed: electrostatic potentials calculation and pKa calculations.
PCE: web tools to compute Protein Continuum Electrostatics.
Nucleic Acids Res. 2005 Jul 1;33(Web Server issue):W372-5.
SPROUTS has been designed to give scientists access to data related to protein folding prediction. In this scope, we processed a set of proteins on five different tools devoted to the prediction of stability changes upon point mutation. We also propose the results obtained with two methods devoted for one to the direct prediction of residues involved in the core of a protein structure and for the other, the characterization of fragments which ends are assumed to be part of the folding nucleus.
SPROUTS: a database for the evaluation of protein stability upon point mutation.
Nucleic Acids Res. 2009 Jan;37(Database issue):D374-9.

TEF is an open-source software for decomposing protein structures into simpler yet informative units named Tightened End Fragments (or closed loops), which can be studied independently to understand protein architecture, folding, and evolution.
TEF2.0: a graph-based method for decomposing protein structures into closed loops.
J Biomol Struct Dyn. 2019 Oct;37(16):4140-4150.
The ProPHet program combines a coarse-grain / elastic network (ENM) protein model and a Brownian Dynamics algorithm to compute protein local rigidity on the residue level.
The program uses a PDB structural file as a starting point and will produce a rigidity profile of the protein under study with a force constant value for each residue in the protein.
Motions and mechanics: investigating conformational transitions in multi-domain proteins with coarse-grain simulations.
Mol. Simul. 2014, 229-236.
