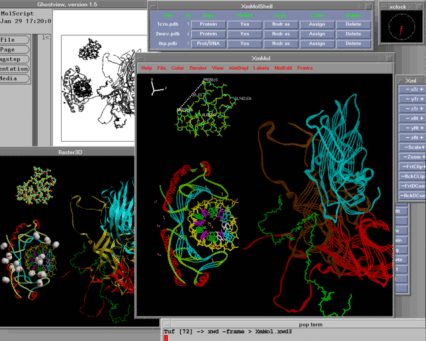
What is XmMol?
XmMol is a desktop macromolecular visualization and modeling tool
designed to be easy to use, configure and enhance. Its graphics are
based on X11, and
part of its user interface is based on Motif. Thus it provides a way of
displaying
structures on any X11 server. Its main features are:
- interactive graphics of macromolecules on any X11 display.
Drawings are performed as wireframes to preserve interactivity. Space
filling static images can be obtained by using an interface to external
rendering programs such as MolScript and Raster3D.
- strong ability to be interfaced with external programs. A
communication protocol allows XmMol to fork external programs called
"delegates" and exchange information. This feature allows the easy
implementation of new features of XmMol. This offers possibilities for
automatic script execution, new file format I/O implementation, file
coordinate modification, implementing external renderers, .... Thus,
XmMol can also be used as a graphic debugger for numerical methods
applied to molecules (minimizers, ...).
Examples of how to implement molecular superimposition, dynamic
trajectory animation, as well as calls to external standard programs
such as babel or hbplus are provided.
- Some modelling tools are supported, such as docking facilities,
interactive backbone deformation (part of the Forme package). However,
the aim of XmMol is mostly to give each user the opportunity to
interface its own methods.
What is XmMol NOT ?
- XmMol does not provide interactive sophisticated space filling
macromolecular representations. Preference was given to portability and
interactivity. Sophisticated space filling representation can however
be obtained through XmMol' interface with MolScript or Raster3D, and
partly with QTree.
- XmMol is not a fully integrated molecular modelling package (see
the above section).

![[Capabilities]](icons/Capabilities.gif)
![[Availability]](icons/Availability.gif)
![[Documentation]](icons/Documentation.gif)
![[Related Packages]](icons/Related.gif)
![[Problems, Help, Feedback]](icons/Help.gif)
Input Formats
PDB and Mac format (XmMol internal format derived from R. Lavery's Flex
format)
are supported as internal formats. Other formats can however be
displayed by using
a converter such as Babel.
Molecular Representations
- Molecules may be drawn as bonds, trace or ribbons. These types
of representations can be assigned to each molecular fragment so that
representations merging all the types can be obtained.
- Various colorations schemes can be applied such as by file,
chain, residue, atom type, secondary structure type, ... The user can
also use a panel to define the color associated with each atom.
- The interface with MolScript and Rasmol was designed to ensure a
good preservation of the view, including representation types as well
as colors. Stereo is preserved with MolScript. A WYSIWYG Postscript
interface is also supported. A mechanism allows, if all necessary
programs are accessible, a direct visualisation of the result.
Display
Any X11 display can be used for visualization (thus works through the
net).
Stereo is supported as two separated images rotated by a few degrees.
Extensibility
XmMol delegate library provides access to any field of the molecular
structures as well
as to most of the XmMol internal functionalities relative to display,
rendering, colouring,
etc. Shared memory usage is supported for some critical fields such as
atomic coordinates.
External delegates can be inserted as part of XmMol menus and made
equivalent to built
in methods.
Configurability
Some parameters can be accessed through a configuration file and
concern:
- Colors used by XmMol, and colors associated with some colouring
patterns.
- Parameters controlling the stereo
- Parameters controlling the spline computation
- Parameters controlling Postscript parameters, and the
possibilities of displaying the result of queries to Raster3D or
MolScript.
- Covalent bond hydrogen bond detection.
- File automatic uncompression.
- Mouse buttons re-assignment.
- Default command file (to make XmMol automatically execute series
of commands)
Hardware requirements
There are no special requirements for XmMol other than Unix and X
windows (X11R4 or upper).
For systems (eg. SUN) that do not support non shared libraries, access
to Motif libraries
should however be ensured.
XmMol was compiled and tested for most unix systems including:
- HP-UX
- SGI IRIX
- IBM AIX
- SUN Solaris
- Dec Alpha
- Linux (should run upon mkLinux)
XmMol is now freely distributed under the terms of the Gnu General
Public License.
To get a copy of the Gnu General Public License, click here.
You can download the distribution
here.
XmMol comes with some "online" documentation as well as with the
PostScript
manuals.
Separately, you can :
- download
the latest PostScript version of the user manual.
- download
the latest PostScript version of the delegate task command language
manual.
For some features (space filling images, molecular superimposition)
XmMol relies on external
programs which are not included in the distribution, or which version
might have been updated.
These include:
None of these libraries/programs are required to run XmMol, but certain
features will not be available without them.
If you need additional information, have a suggestion, or need help
concerning XmMol, please first read the documentation, including some
Frequently Asked Questions. You can also contact the team at RPBS.

Last modified: Mon Jan 27 18:38:11 2008
![]()