We use the same output page to display results of mdpocket
job for both server. Before describing it, we just describe
for both servers, how to reach this page. Don't worry it's
really easy.
Getting to the results page
Default server
Just follow the
input tutorial,
where we describe all intermediates pages from which you
will be able to reach the final results page.
Advanced server (Mobyle)
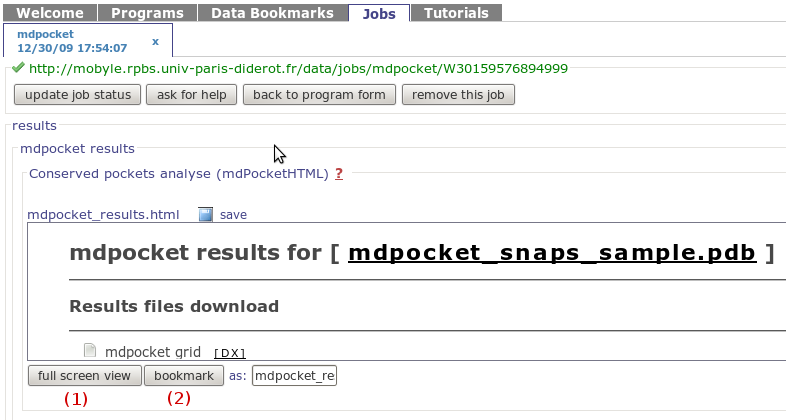
In mobyle, once your job will be finished, you will be
redirected to the mobyle results page. This results page
is shown in the snapshot on the right. The results page
is actually contained in the "Conserved pockets analyse
alpha spheres (mdPocket HTML)" labeled scroll pane. To have
a more friendly view and reach the output pages described
just click on the Full screen view
button (1)
You also have the possibility to bookmark the result page
using the Bookmark button
(2)
(enter the bookmark name in the field on left)
Main program output results are provided both in the final
results page and in the mobyle interface. This redundancy
is comfortable to (i) analyse your results in a specific
and integrated web page and (ii) to directly use the
mobyle pipelining feature for further analysis.
Results page description: basic mdpocket run (mode 1)
The results can be roughly divided in 3 sections:
Output files,
Snapshots and
Visualisation .
Output files
Several mdpocket output files are provided and described
below. You can download all of them.

-
MDpocket grid
(1): it is the MDpocket
output grid that stores density information for each
grid point. To learn more about mdpocket methodology
and output, go tho the
mdpocket method section
in the homepage, or read
the documentation.
-
Pocket grid points
(2): This file contains
all grid points having 3 or more Voronoi Vertices in
the 8A3 volume around the grid point for each snapshot.
This output can be used to define a specific zone (a pocket)
on which you may want to make further analysis using mdpocket
round 2.
Additionally, we provide 3 pre-defined PDB files containing
grid points up to a given threeshold. Low, medium and high
resolution corresponds to low, medium and high isovalues, and
thus can be seen as the grid points corresponding to all
conserved, and highly conserved cavities.
-
Pocket density
(3): This file contains
the first snapshot, written as PDB file, with the
B-factor value matching alpha spheres density nearby this atom.
This file allows intuitive, coloured visualisation
similar to that of the snapshots described below.
<Back to top>
Snapshots
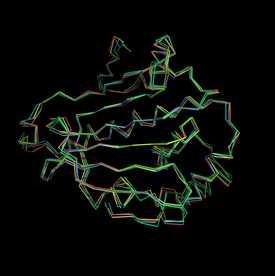
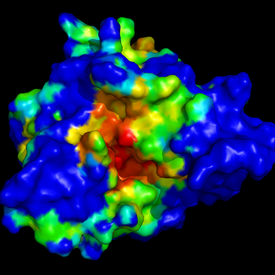
Two sets of snapshots are provided here: the first set
(left picture) represents an alignement of all input
structures, and the second set (right picture) represents
the first structures surface coloured by alpha spheres
density. Here, coulours range from blue (low density =
no particular cavity at this place) to red (high density
= hot spot = conserved cavity!).
Note that you may obtain such a display by dowloading the output
PDB file "Pocket density" described previously, and display
it using PyMOL and VMD (color the molecular surface by B-factor).
Hopefully, we will allow you to do so directly from here
with a new interactive viewer making use of OpenAstex viewer.
<Back to top>
Visualisation
Currently, the visualisation is made using both Jmol and
OpenAstex, in which the mdpocket output PDB file is
automatically loaded in each viewer
(1) (first snapshot).
Using Jmol, you can view the Grid file
extracted from mdpocket results. A slider is provided to
change isovalue: the highest the isovalue is, the more
conserved is the corresponding cavity.
Using OpenAstex, the visualisation is
atom-centered. That is, the isovalues have been mapped
from the grid to the atoms and transformed to be somehow
B-factor-like for coloring purposes. The highest the
B-factor is, the more conserved is the cavity associated
with atoms.
Both visualisation methods use different metrics.
Isovalues will tipically range from 0 to N, with N having no real
limitation (depends on the number of snapshots; the slider
is limited to 800), while B-factor will range from 0 to 7-8,
as it is log-scaled. We are investingating a way to get
a common metric, and to merge these visualisation features
in a single viewer.
Jmol
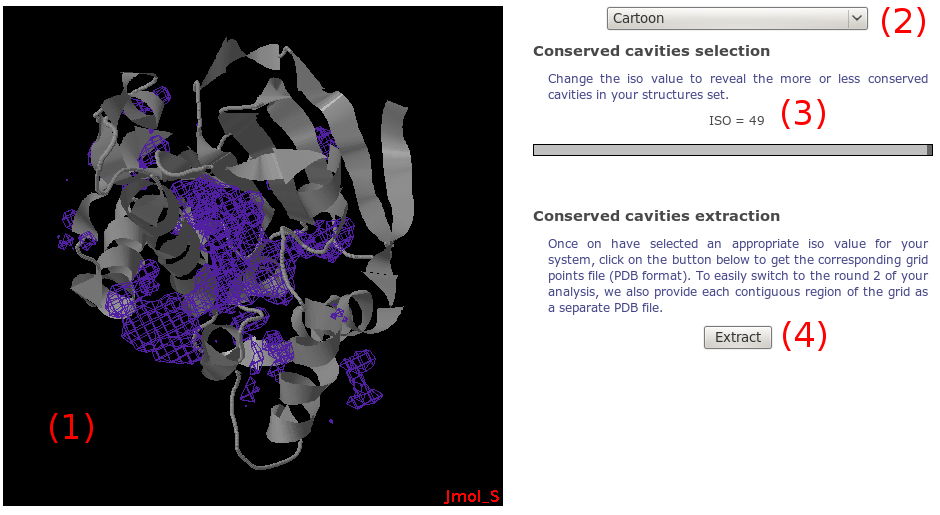
Using Jmol, the density grid is loaded along with the protein
(1).
On the right, you have a set of graphic components to
facilitate the viewing. A simple selection box
(2) allows you to perform
basic changes to the whole system representation
(display protein as cartoon, reset view...).
Using the slider on the right, you can change
(3) the so called "isovalue".
For a given grid point, this isovalue represents the number
of alpha spheres seen for all snapshots within a 8A radius.
Thus, a high isovalue will display protein cavities which are
conserved during the MD, while low isovalues will rather show
you every potential transient pockets.
Finally, we give you the opportunity to downlad grid points
(PDB format) corresponding to each contigous points that
could form a potential pocket(4).
To do so, we use an internal clustering procedure from Pymol.
You can then use these files as input for the second round of
mdpocket (see inputs and ouptuts
description), to monitor a specific part of the protein,
basically a (potential?) binding site of interest.
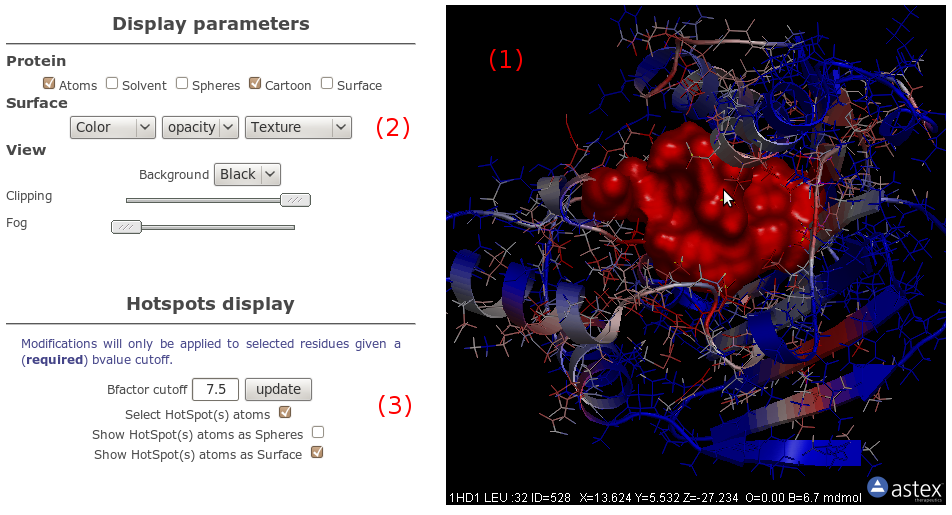
OpenAstexViewer
Using OpenAstex, atoms are coloured by B-factor, with the same
convention as that used for snapshots, that is, the more an
atom is close to red (resp. blue), the more conserved is
its corresponding cavity. Only here, the grid isovalue have
been mapped on atoms (see accompagning paper for formula)
so you can see directely which atom is associated with a
conserved zone.
Besides basic visualisation options (2),
we give you the possibility to select atoms based on B-factor
value (3). By checking the
appropriate checkbox, all atoms having B-factor up to the
specific value (in the text field) will be selected.
If you change this isovalue cutoff, you have to click the
update button to update atom selection.
0 is the minimum B-factor value, while the maximum should
lie around 7 or 8 (due to log-scaling of isovalues).
Remember that if you know Jmol and OpenAstex, you can
access the display popup menu by right clicking on the view.
Results page description: pocket analysis
The pocket analysis will provide 3 output files. Note
that there is currently no specific interactive visualisation
in the result page, we just provide the result files as
is (like in the future mdpocket distribution).

-
Pocket vertices
(1):
This is a pdb file that contains all Voronoi vertices
in the selected pocket zone for each snapshot. Each
snapshot is handled as separated model (like a NMR
structure) and can thus be viewed as MD using PyMOL.
Show the surface of the vertices and you can visualize
the movement of your pocket. Be careful, VMD does not
read this file, as from one snapshot to the other a
different number and type of Voronoi vertices can be
part of the model.
-
Pocket atoms
(2):
This is a pdb file similar to the previous output, but
this time containing all receptor atoms defining the
binding pocket.
-
Pocket descriptors
(3):
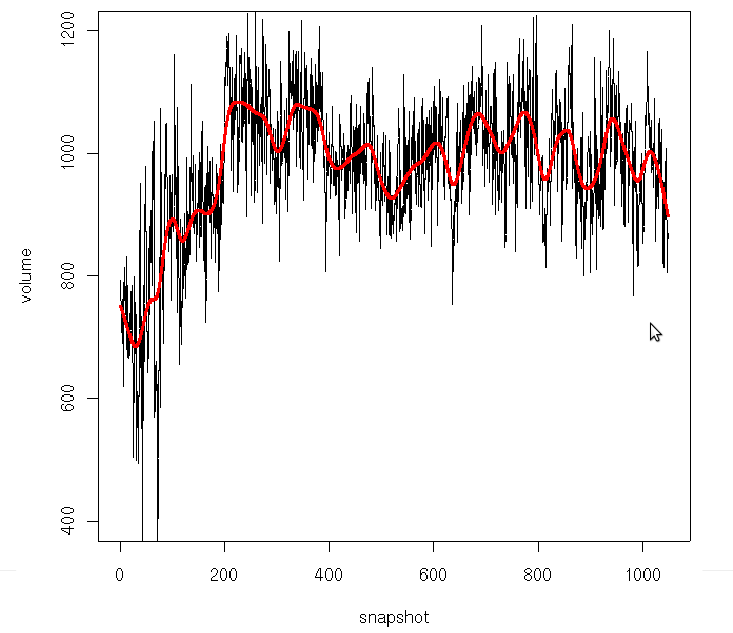
This file contains all pocket descriptors calculated
by MDpocket for each of the input snapshots (1 per line).
You can therefore see the evolution of these descriptors,
for example the pocket volume...
As this output file is a simple text file, it can be
easily analyzed using R, gnuplot or other suitable
software. An example R output for the pocket volume would be: